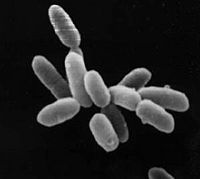
Halobacterium NRC-1
For the course duration, the article is closed to outside editing. Of course you can always leave comments on the discussion page. The anticipated date of course completion is May 21, 2009. One month after that date at the latest, this notice shall be removed. Besides, many other Citizendium articles welcome your collaboration! |
| Halobacterium sp. NRC-1 | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | ||||||||||||||
| Scientific classification | ||||||||||||||
| ||||||||||||||
| Binomial name | ||||||||||||||
| Halobacterium sp. NRC-1 |
Description and significance
Halobacterium sp. NRC-1 is a halophilic archaea which thrives all over the world in high salt environments, including salt production facilities, brine inclusions in salt crystals, natural lakes and ponds, and salt marshes. Prior to 1990 H. NRC-1 was classified as an archeabacterium under the prokaryote kingdom in the two-empire system which consisted of eukaryotes and prokaryotes. Since 1990 the prokaryotes were split into bacteria and archaea due to their different evolutionary paths and biochemistry differences.[1] Like all archaea H. NRC-1 has no nucleus or organelles within the cell, and like other archaea, have evolved many metabolic pathways to allow it to survive in extreme environments.[2] Halobacterium sp. NRC-1 is motile using both flagella and gas vesicles, and respond to their environment by moving towards chemicals using a process called chemotaxis and toward or away from light using phototaxis using its sensory rhodopsins. They reproduce via binary fission and grow best in a 42 degree Celsius aerobic high salt environment.
Halobacterium sp. NRC-1 is very easy to culture in the lab and its genome has been completely mapped. Whole-genome DNA microarrays are available to investigate gene expression. This makes it an excellent model microorganism for research into the basic cellular process and gene expression as well as for teaching.
Genome structure
The genome of Halobacterium sp. NRC-1 was published in 2000. Since that time scientists have used a combination of genetic, transcriptomic, proteomic and bioinformatic approaches to gain insights into both its extremophilic lifestyle as well as the fundamental cellular processes common to all life forms.[3] In order to survive in its extreme environment H. NRC-1 has developed extensive capabilites to repair its own genome. In combination with its saline environment which provides some protection from UV radiation, its ability to repair its chromosomes after extensive damage is only exceeded by the extremely radiation resistant Deinococcus radiodurans.[4][5]
- Halobacterium sp. NRC-1 contains the smallest genome to date among the halophiles. It is 2,571,010 bp in size, and is composed of a large GC-rich chromosome (2,014,239 bp, 68 % G+C), and two smaller extrachromosomal replicons, pNRC100 (191,346 bp) and pNRC200 (365,425 bp), with 58–59 % G+C composition. The two smaller replicons contain 145,428 bp of identical DNA and 33–39 kb inverted repeats catalyzing inversion isomers, and the majority of the 91 IS elements, representing 12 families, found in the genome. As a result of the large number of repeated sequences, genome assembly required extensive genomic mapping and an ordered clone library of pNRC100. Of the 2,630 likely protein-coding genes in the genome, 2,532 are unique. Halobacterium predicted proteins were found to be highly acidic and a substantial number had bacterial homologs as their closest relatives, suggesting that they might have been acquired through lateral gene transfer. In addition, 52 RNA genes were also identified; however, the 16S rRNA sequence and other unique characteristics did not allow placement within a validly described Halobacterium species, and this point has been the subject of some controversy. Interestingly, about 40 genes in pNRC100 and pNRC200 code for functions likely to be essential or important for cell viability (e.g. thioredoxin and thioredoxin reductase, a cytochrome oxidase, a DNA polymerase, multiple TATA-binding proteins (TBP) and transcription factor B (TFB) transcription factors, and the only arginyl-tRNA synthetase in the genome).[3]
While much of the published literature on this organism refers to these smaller genetic elements as replicons or megaplasmids,they don't really fit the definition of these terms. Replicons are considered to be exact copies of specific sequences of an original DNA or RNA genome, or even a whole copy of the original genome. Plasmids are referred to as being small extra chromosomal DNA elements that carry relatively few genes that code for genetic information that is not essential to an organism's biological processes. Considering that the genetic information carried on these replicons not only code for information that is essential to the organism's survival, but also contain nucleotide sequences that are not identical to that of it's larger chromosome, scientists are beginning to refer to them as "minichromosomes" rather than "megaplasmids".
Cell structure and metabolism
Halobacterium sp. NRC-1 is an obligate halophilic microorganism which has adapted to be able to grow in conditions of extremely high salinity, up to 10 times that of seawater.[6] In order to survive under these conditions it maintains a very high concentration of salts internally in the form of KCl to enable it to remain isotonic to it's preferred environment.[7] Halorhodopsin plays a very energy efficient role in helping to maintain the osmotic balance within the cell by acting as a light driven pump to transport clorine ions into the cell.[8] This microorganism has been extensively studied and shown to contain some of the classic features found in halophilic archaea, for example, an S-layer glycoprotein, ether-linked lipids, and purple membrane.[9]
The purple membrane consists of the light-driven ion transporters bacteriorhodopsin and halorhodopsin, and the phototaxis receptors, sensory rhodopsins I and II.[6] In order to survive in low oxygen environments, Halobacterium sp. NRC-1 synthesizes Bacteriorhodopsin, which is a unique protein that can use light as an energy source, much like chlorophyll can in cyanobacteria and phototrophic eukaryotes. When the retinal in in Bacteriorhodopsin absorbs light, it results in a series of conformational changes that translocates the proton into the periplasmic space. This light driven proton pumping generates a pH gradient which is then used to power the synthesis of ATP by chemiosmosis. This phototrophic capability is particularly useful to Halobacterium sp. NRC-1 as oxygen is not very soluble in concentrated salt solutions. In addition to its phototrophic respiration capabilities, is also capable of anaerobic respiration using dimethyl sulfoxide (DMSO) and trimethylamine-N-oxide (TMAO).[10]
- Halobacterium NRC-1 is an aerobic chemoorganotroph, growing on the degradation products of less halophilic organisms as the salinity reaches near saturation. In the laboratory, cells are cultured best in a complex medium. A minimal medium described for Halobacterium includes all but 5 of the 20 amino acids for growth. Several amino acids may be used as a source of energy, including arginine and aspartate, which are passed to the citric acid cycle via 2-oxoglutarate and oxaloacetate, respectively. Under aerobic conditions, arginine is presumably converted to glutamate via the arginine deiminase pathway, and this amino acid then enters the cycle via glutamate dehydrogenase. The arginine deiminase pathway is coded by the arcRACB genes, which are found on pNRC200.[6]
Ecology
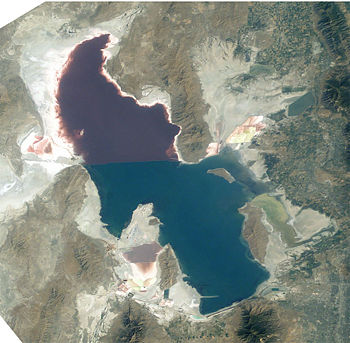
Halobacterium sp. NRC-1 is one of many strains of halobacterium which thrive in extremely high salinity environments such as salt lakes, salt marshes and salt drying ponds. Their optimal temperature for reproduction is 42°C. Often these highly saline bodies of water will be tinted red or purple. It is the red/purple color of the bacteriorhodopsins that give the red color you often see in these highly saline environments. Bacteriorhodopsin consists of a photosensitive pigment protein called retinal. This protein pigment is responsible for NRC-1's colorful properties. The more saline the environment the redder the color will be because halobacterium increase their production of bacteriorhodopsin in response to drops in oxygen which is less soluable in saline solutions. There are not many other organisms that can survive in these high salt environments, in fact one of its primary sources of food is the amino acids of other organisms which have lysed due to the high salt concentration in this environment. Brine shrimp are one of a few other organisms that can survive the high salt concentration, and they feed almost exclusively on the bacteria in their environment. Interestingly, the flamingo, whose pink color comes exclusively from its diet (it doesn't have the ability to make the carotinoids which give the pink color) feeds on the brine shrimp, so the carotinoids which give the flamingo its pink color actually come from the rhodopsins in the halobacterium that the shrimp eat and whatever carotinoids that the shrimp produce naturally.
Application to Biotechnology
Halobacterium sp. NRC-1 was one of the first Archaea to have its genome fully mapped and published.[6] Since that time several more species have been successfully mapped and a few others partially mapped. This allows scientists to analyze haloarchaeal properties in silico to determine the activity of genes. In addition many transformation tools have been developed to allow the isolation of genes necessary for biological processes by complementation of loss-of-function mutants.[11] This makes it an ideal model for testing the function of genes.
In addition to its usefulness as a genetic test bed, Halobacterium sp. NRC-1 is an ideal study tool because it is an extremophile, it exists in environments that are very unfriendly toward life. This makes it very useful for studying many biological questions especially those involved with adaptation and survival in extreme environments. In fact, much research is being done to investigate whether or not halobacterium could be potential candidates for extraterrestrial life, such as on Mars or Europa. Recent research has shown that Halobacterium sp. NRC-1 can not only survive at temperatures far below its optimal growth temperature, but continue to reproduce as well.[12]
Current Research
Survival and growth of Halobacterium sp. NRC-1 following incubation at -15°C, freezing or freezedrying, and the protective effect of cations
Since extraterrestrial halite has been discovered in meteorites from Mars and since several extremely halophilic archaea were isolated from geologically ancient rock salt, the behaviour of halophilic microorganisms in Martian conditions is being investigated. Low temperatures and low water activity are particularly characteristic of the Martian surface. Experimental procedures were developed for testing the response of halobacteria to Martian or other extreme environmental conditions.[12]
Functional Genomics of Thioredoxins in Halobacterium sp. NRC-1
This project addresses the functions of an ancient protein family in Archaea that occupy extreme environments. Some of these proteins may play roles similar to those of comparable proteins in other living organisms, and thus may tell us about functions that evolved in the last universal common ancestor of life. Others may have evolved as the Archaea began to occupy specialized and often extreme environments. This project also addresses the emergence of proto-metabolic networks that supplied the precursors for the RNA World.[13]
A systems view of haloarchaeal strategies to withstand stress from transition metals
Given that transition metals are essential cofactors in central biological processes, misallocation of the wrong metal ion to a metalloprotein can have resounding and often detrimental effects on diverse aspects of cellular physiology. Therefore, in an attempt to characterize unique and shared responses to chemically similar metals, we have reconstructed physiological behaviors of Halobacterium NRC-1, an archaeal halophile, in sublethal levels of Mn(II), Fe(II), Co(II), Ni(II), Cu(II), and Zn(II).[14]
References
- ↑ Woese, C R (1994-03). "There must be a prokaryote somewhere: microbiology's search for itself". Microbiological Reviews 58 (1): 1-9. ISSN 0146-0749. Retrieved on 2009-04-28.
- ↑ Archaea - Wikipedia, the free encyclopedia. Retrieved on 2009-04-26.
- ↑ 3.0 3.1 DasSarma S et al. (2006), "Post-genomics of the model haloarchaeon Halobacterium sp. NRC-1", Saline Systems 2 (3), DOI:10.1186/1746-1448-2-3
- ↑ McCready, Shirley; Jochen Muller, Ivan Boubriak, Brian Berquist, Wooi Ng, Shiladitya DasSarma (2005). "UV irradiation induces homologous recombination genes in the model archaeon, Halobacterium sp. NRC-1". Saline Systems 1 (1): 3. DOI:10.1186/1746-1448-1-3. ISSN 1746-1448. Retrieved on 2009-04-18. Research Blogging.
- ↑ Kottemann, Molly; Adrienne Kish, Chika Iloanusi, Sarah Bjork, Jocelyne DiRuggiero (2005-06-01). "Physiological responses of the halophilic archaeon Halobacterium sp. strain NRC1 to desiccation and gamma irradiation". Extremophiles 9 (3): 219-227. DOI:10.1007/s00792-005-0437-4. Retrieved on 2009-04-28. Research Blogging.
- ↑ 6.0 6.1 6.2 6.3 Ng, Wailap Victor, et al. (2000-10-24). "Genome sequence of Halobacterium species NRC-1". Proceedings of the National Academy of Sciences of the United States of America 97 (22): 12176-12181. DOI:- 97 VL - 97. Retrieved on 2009-04-18. - 97 Research Blogging.
- ↑ Coker, James; Priya DasSarma, Jeffrey Kumar, Jochen Muller, Shiladitya DasSarma (2007). "Transcriptional profiling of the model Archaeon Halobacterium sp. NRC-1: responses to changes in salinity and temperature". Saline Systems 3 (1): 6. DOI:10.1186/1746-1448-3-6. ISSN 1746-1448. Retrieved on 2009-04-18. Research Blogging.
- ↑ Oesterhelt, D (1998-08). "The structure and mechanism of the family of retinal proteins from halophilic archaea". Current Opinion in Structural Biology 8 (4): 489-500. ISSN 0959-440X. Retrieved on 2009-04-26.
- ↑ Kennedy, S P; W V Ng, S L Salzberg, L Hood, S DasSarma (2001-10). "Understanding the adaptation of Halobacterium species NRC-1 to its extreme environment through computational analysis of its genome sequence". Genome Research 11 (10): 1641-50. DOI:10.1101/gr.190201. ISSN 1088-9051. Retrieved on 2009-04-18. Research Blogging.
- ↑ Müller, Jochen A.; Shiladitya DasSarma (2005-03). "Genomic Analysis of Anaerobic Respiration in the Archaeon Halobacterium sp. Strain NRC-1: Dimethyl Sulfoxide and Trimethylamine N-Oxide as Terminal Electron Acceptors". Journal of Bacteriology 187 (5): 1659–1667. DOI:10.1128/JB.187.5.1659-1667.2005. Retrieved on 2009-04-18. Research Blogging.
- ↑ Soppa, Jorg (2006-03-01). "From genomes to function: haloarchaea as model organisms". Microbiology 152 (3): 585-590. DOI:10.1099/mic.0.28504-0. Retrieved on 2009-04-18. Research Blogging.
- ↑ 12.0 12.1 Weidler, Gerhard (2004-03-01), Survival and growth of Halobacterium sp. NRC-1 following incubation at -15°C, freezing or freeze-drying, and the protective effect of cations, Third European Workshop on Exo-Astrobiology, vol. 545, at 311-312. Retrieved on 2009-04-18
- ↑ Library of Resources « NASA Astrobiology. Retrieved on 2009-04-18.
- ↑ Kaur, Amardeep; Min Pan, Megan Meislin, Marc T. Facciotti, Raafat El-Gewely, Nitin S. Baliga (2006-07). "A systems view of haloarchaeal strategies to withstand stress from transition metals". Genome Research 16 (7): 841–854. DOI:10.1101/gr.5189606. Retrieved on 2009-04-18. Research Blogging.