Staphylococcus epidermidis
| Staphylococcus epidermidis | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | ||||||||||||||
| Scientific classification | ||||||||||||||
| ||||||||||||||
| Binomial name | ||||||||||||||
| Staphylococcus epidermidis |
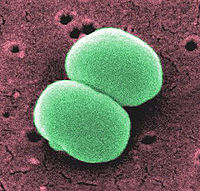
Staphylococcus epidermidis is a Gram-positive coccus, nonpigmented, bacterium. This organism, usually 0.5 to 1.5 mm in diameter, is a normal inhabitant of the human skin that grows in clusters. Research studies reveal that S. epidermidis lives in close association with S. aureus, a very destructive pathogen.
Even though a coagulase-negative Gram bacterium, S. epidermidis has been lately classified among the most important pathogens responsible for diverse nosocomial infections. Most strains are highly resistant to multiple antibiotics, such as penicillin, tetracycline, methicillin and many more, which makes it very difficult to treat the infections resulting from these bacteria. The Centers for Disease Control and Prevention's National Nosocomial infection surveillance system report that S. epidermidis is responsible for 33.5% of nosocomial blood stream infections[1].
Genome structure
Among all the S. epidermidis strains, the genome sequence of only 2 strains has been completely described: S. epidermidis RP62a with a genome length of 2,616,530 bp and S. epidermids ATCC12228, 2,499,279 bp. RP62a encodes 2585 protein genes, 61 tRNA and 19 rRNA; whereas the ATCC12228 strain contains a number of 2381 protein coding genes, 60tRNA and 16rRNA. The genome of S. epidermidis has a low G+C content (32.1%), which is believed to contribute to the virulence and the high resistance of this strain against many antimicrobial agents. The genomic elements, which include, the genome islands, the insertions sequences, the composite transposons and the integrated plasmids, constitute more or less 9% of the genome[2].
Studies have shown that a high level of genetic diversity exists within S. epidermidis at the level of a single locus as a result of adaption to different environment in hospitals and communities[3].
Cell structure and metabolism
As a Gram-positive bacterium, S. epidermidis has a cell wall made of a large concentration of peptidoglycan layer but no outer membrane. The cell-wall anchored(CWA)proteins of these bacteria are part of a family of surface-exposed proteins that interact with targets in the host. These CWA proteins possess an N-terminal secretion signal sequence (S) required for sec-dependent secretion, followed by a non-repeat A domain that contains the ligand-binding site[4] . These interactions are particular important because they are involved in bacterial adherence and escape from the host defense systems. No mechanism is known to explain the virulence of S. epidermidis, but it is thought that the cell wall proteins are potential factors that must be involved.
S.epidermidis requires mannitol as the carbon source to grow in aerobic conditions; however, this organism also survives anaerobically when placed in a standardized complex medium of glucose[5] . It is non-motile and has no endospores. Since it is a pathogen of the human skin, S. epidermidis grows best at 37°C (mesophilic), the optimal temperature for the human body.
Pathology
Most infections caused by S. epidermidis affect hospital patients because these organisms develop structures called biofilms on the surface of implanted medical devices. These biofilms result from the agglomeration of bacterial cells that are contained in an exopolysaccharide matrix, the slime, to produce a very dense and protected environment from the host defense systems and antibiotics. They not only provide protection from the host but they also result in the damage of surrounding tissues of the host.[6]
Skin and tissues infections, post-surgical wounds are the common points of entry for infection by S. epidermidis. This bacterium accounts for most catheter-related infections, joint replacement infections, the majority of prosthetic cardiac valve infections and infections following post-neurosurgical procedures.In addition, studies have reported that S. epidermidis is highly predominant in the milk and feces of breast-fed infants compared to formula-fed ones. However, whether or not the presence of these bacteria can cause infections in these children have not been determined.
Moreover, S. epidermidis plays a significant role in some external ocular diseases, such as chronic blepharitis and suppurative keratitis. Fortunately, these infections are treatable; and ciprofloxacin is considered the best drug in the cure of bacterial keratitis. Interestingly, methicillin-resistant S. epidermidis was identified in the cause of nosocomial meningitis when prosthetic devices were used in a trauma case in 2003. Even though, the antibiotic, vancomycin, was the treatment of choice, it was not successful in the recovery of the patient. Instead, better results were obtained with the use of linezolid[7] . Due to their high resistance to antimicrobial agents, other techniques have been employed to prevent the staphylococcal infections which imply mechanical cleaning, the stopping of biofilm growth by the removal of essential nutrients, the inhibition of microbial attachments to surfaces, and biomass detachment[8].
Application to Biotechnology
The pulse-field gel electrophoresis is the most common method used to study methicillin-resistant S.epidermidis. It allows scientists to determine the cause of nosocomial infections and the means of transmission of this bacterium. Because S. epidermidis does not release toxins, it is very difficult to trace the infections, but some molecular genetics technologies, such as the automated culture system BacT/ALERT™ )[9] in Canada and the Real-Time Reverse Transcriptase Polymerase Chain Reaction (PCR) Assay, nhave been employed in the testing of bacterial contamination of blood platelets. Moreover, multilocus sequence typing (MLST)[10] is implemented to study the phylogenetic relationships of divers bacterial pathogens, including S. epidermidis and S. aureus.
Current Research
Two decades ago, S. epidermidis was considered a nonsignificant, simple Gram-positive bacterium. However, there has been lately a growing interest in the study of this organism due to its high level of pathogenicity in nosocomial infections. A lot of research is in progress at Baylor College of Medicine in the determination of the genome sequence of different strains, such as S. epidermidis M23864:W2, S. epidermidis CONS. Besides, more studies are currently implemented so as to reveal the mechanisms of virulence and transmission at the molecular and cellular level. There is also a great focus on the research of more effective preventive inventions agaisnt these infections. Because S. epidermidis is highly resistant to most antibiotics and lightly resistant to only vancomycin, it is very difficult to treat the nosocomial infections. But a class of antimicrobial agents, the cationic antimicrobial peptides (AMPs), are actually under study,which are thought to kill the microbial cell by pore formation or cell disruption.
Enzymatic Detachment of Biofilm
The formation of biofilms by S. epidermidis has been accounted for the highly resistance of these pathogens against a large number of antibiotics. Consequently, it has become a very difficult task to treat the nosocomial infections caused by S.epidermidis. In this experiment, researchers have tested the effects of an enzyme, called dispersin B, on the possible detachment of the biofilms from the surfaces of medical devices. Dispersin B is biofilm enzyme released by the Gram-negative pathogen Actinobacillus actinomycetemcomitans. In the course of this experiment, researchers grew 4 strains of S. epidermidis from infected intravenous catheters by serial dilution on the wells on a 96-well polystyrene microtiter plate. After a few hours of incubation, the amount of biofilm produced was evaluated by measuring the absorbance of the wells at 590 nm using a microtiter plate reader. However, when the cells were washed with dispersin B, the results yielded from the absorbance reading revealed that very little or no biofilms grew on the wells. Hereby, dispersin B represents now a possible weapon in the treatment of nosocomial infections[11].
Intercellular Adhesin Production Significantly Increases
Instead of using antimicrobial agents, another group of researchers have studied the effects of altering TCA cycle activity on the development of biofilm. The study was conducted by using fluorocitrate, a TCA cycle inhibitor. The research revealed that decreasing TCA activity by fluorocitrate would cause an increase in the concentration of polysaccharide intracellular adhesin (PIA), an important protein necessary for the development of biofilm. Besides fluorocitrate, any other factor, such as low level of oxygen, lack of iron, affecting TCA activity would increase the level of PIA. As a result, researchers have concluded that increasing TCA cycle activity would cause a decrease in PIA production, thus triggering removal of biofilms on the surface of medical devices. It is now being investigating the probability of altering PIA production by altering the availability of NADH or NAD+ in the environment[12].
Protection with Human Immunoglobulin G
The study was conducted in the purpose of testing the effects of increasing the levels of INH-A21, which is an intravenous immunoglobulin from donors with high titers of antistaphylococcal antibodies, on both S. epidermidis and S. aureus. The main focus of the study was to determine the extent to which this increase in INH-A21 would protect very low-birth weight (VLBW) infants and infected patients against nosocomial infections produced by these strains. INH-A21 works by recognizing and inhibiting fibrinogen binding by ClfA and SdrG, which would reduce the capacity of staphylococci to spread hematogenously and bind to its sites of vascular damage. Opposite results were obtained on the effects of INH-A21 in VLBW infants: a group of researchers found that this protein does protect against staphylococcal infections whereas another group found that INH-A21 does not reduce the level of occurrency of nosocomial infections in very low birth weight infants. Nevertheless, it has been proven that INH-A21 does protect infected patients other than infants against staphylococcal infections, and may eventually become an important tool in the treatment of these infections, but it is still not known why the effects were controversal in VLBW infants[13]
References
- ↑ Rasheed, M. U (2007). "Staphylococcus epidermidis: A Commensal Emerging As A Pathogen With Increasing Clinical Significance Especially In Nosocomial Infections". the internet journal of Microbiology 3 (2): 1453-1464. Retrieved on 2009-04-20.
- ↑ Gill, Steven R. (2005). "Insights on Evolution of Virulence and Resistance from the Complete Genome Analysis of an Early Methicillin-Resistant Staphylococcus aureus Strain and a Biofilm-Producing Methicillin-Resistant Staphylococcus epidermidis Strain". American Society for Microbiology 187 (7): 2426-2483. DOI:10.1128/JB.187.7.2426-2438.2005.. Retrieved on 2009-04-20. Research Blogging.
- ↑ Miragaia, M (2007). "Inferring a Population Structure for Staphylococcus epidermidis from Multilocus Sequence Typing Data". Journal of Bacteriology 189 (6): 2540-2552. Retrieved on 2009-04-20.
- ↑ Bowden, Gabriela M. (2005). "Identification and preliminary characterization of cell-wall-anchored proteins of Staphylococcus epidermidis". Microbiology 151: 1453-1464. DOI:10.1099/mic.0.27534-0.. Retrieved on 2009-04-20. Research Blogging.
- ↑ Schaefler, S. (1971). "Staphylococcus epidermidis BV: Antibiotics Resistance Patterns, Physiological Characteristics, and Bacteriophage Susceptibility". American Society for Microbiology 22 (4): 693-699. Retrieved on 2009-05-08.
- ↑ O'Gara, James P. (2001). "Staphylococcus epidermidis biofilms: importance and implications". Journal Medicine of Microbiology 50: 582-587. ISSN 0022-2615. Retrieved on 2009-04-20.
- ↑ Krueger, Wolfgang A. (2004). "Treatment of Meningitis Due to Methicillin-Resistant Staphylococcus epidermidis with Linezolid". Journal of Clinical Microbiology 42 (2): 929-932. DOI:10.1128/JCM.42.2.929-932.2004.. Retrieved on 2009-04-21. Research Blogging.
- ↑ Joao, Xavier (2005). "Biofilm-control strategies based on enzymic disruption of the extracellular polymeric substance matrix – a modelling study". Microbiology 151: 3817-3832. DOI:10.1099/mic.0.28165-0. Retrieved on 2009-04-20. Research Blogging.
- ↑ Ramirez-Arcos, Sandra M. (2007). "Method for detection of Staphylococcus epidermidis description/claims". Journal of Bacteriology. Retrieved on 2009-04-21.
- ↑ Miragaia, M (2007). "Inferring a Population Structure for Staphylococcus epidermidis from Multilocus Sequence Typing Data". Journal of Bacteriology 189 (6): 2540-2552. Retrieved on 2009-04-20.
- ↑ Kaplan, Jeffrey B. (2004). "Enzymatic Detachment of Staphylococcus epidermidis Biofilms". Antimicrob Agents Chemother 48 (7): 2633-2636. DOI:10.1128/AAC.48.7.2633-2636.2004.. Retrieved on 2009-05-08. Research Blogging.
- ↑ Vuong, Cuong (2005). "Staphylococcus epidermidis Polysaccharide Intercellular Adhesin Production Significantly Increases during Tricarboxylic Acid Cycle Stress". Journal of Microbiology 187 (9): 2967-2973. DOI:10.1128/JB.187.9.2967-2973.2005.. Retrieved on 2009-05-09. Research Blogging.
- ↑ Vernachio, John H. (2006). "Human Immunoglobulin G Recognizing Fibrinogen-Binding Surface Proteins Is Protective against both Staphylococcus aureus and Staphylococcus epidermidis Infections In Vivo". Antimicrobial Agents and Chemotherapy 50 (2): 511-518. DOI:10.1128/AAC.50.2.511-518.2006.. Retrieved on 2009-05-13. Research Blogging.